| PyMOL |
 |
Drag the .pml script from Explorer/Finder/Dolphin etc. into PyMOL window.
Using a terminal (e.g. xterm):
# run a PyMOL command script (opens PyMOL GUI) pymol script.pml # run a Python script pymol script.py # batch mode (no PyMOL GUI) pymol -c script.pml # use absolute path if "pymol" is not in $PATH /opt/pymol-2.2.3/pymol -c script.pml
alias pymol=/Applications/PyMOL.app/Contents/MacOS/PyMOL
Register .pml extension with PyMOLWin.exe
Using Windows Explorer:
Load PML scripts with @:
PyMOL> @example.pml
PyMOL> @/tmp/PyMOLScripts/example.pml
PyMOL> @C:\PyMOLScripts\example.pml
Load Python scripts with run:
PyMOL> run example.py
PyMOL> run /tmp/PyMOLScripts/example.py
PyMOL> run C:\PyMOLScripts\example.py
Download the PyMOL (PML) script example.pml and open it with PyMOL by dragging it into the PyMOL window.
Expected result:
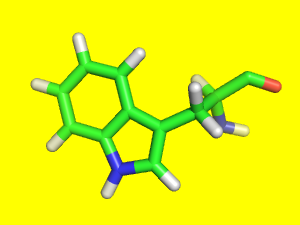
Run the same example.pml script on the PyMOL command line with @.
Hints: The commands cd, pwd and ls are useful to first navigate to the right directory. Filename auto-completion speeds up typing and avoids typos. You can also use environemnt variables (e.g. $HOME) and ~ (home directory).
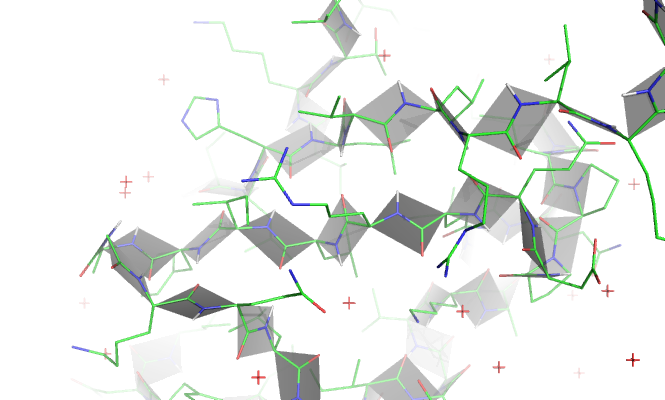
run ~/Downloads/bbPlane.py
fetch 1ubq
bbPlane
Expected result:
© 2017 Schrödinger, Inc.